Overview. Hospitals today are collecting an immense amount of patient data (e.g., images, lab tests, vital sign measurements), but are still ignoring the vast majority of it. Despite the fact that health data are messy and often incomplete, these data are useful and can help improve patient care. To this end, we have pioneered work in leveraging machine learning (ML) and electronic health records for predicting adverse outcomes or events (e.g., infections). Based on collaborations with 30+ clinicians, we have identified key characteristics for the safe and meaningful adoption of ML in healthcare. Beyond accuracy, models must be, actionable (tell a clinician how to reduce a patient’s risk not just who’s at risk) and robust (capable of adapting to changes across populations and time). Achieving accurate models with these characteristics presents unique technical challenges. Specifically, in healthcare, one often deals with high-dimensional data (i.e., many covariates) but has few examples to learn from –‘high D small N.’ To address these challenges, we have developed new ML techniques.
Methodological Contributions
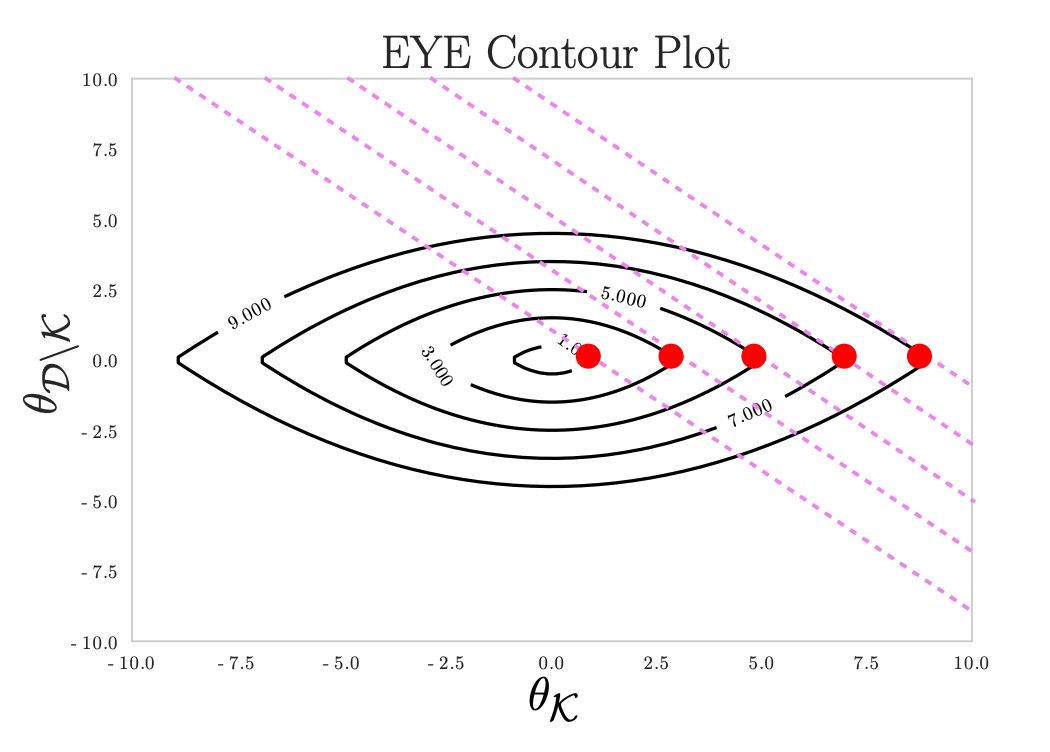
Incorporating Domain Expertise – recently, we proposed a new regularization penalty, the EYE penalty, that incorporates domain knowledge regarding known risk factors. In ML, and specifically deep learning, domain knowledge is oftentimes ignored. However, in healthcare, domain knowledge can be critical when training data are limited. Our proposed approach uses domain knowledge to help select among highly correlated variables. This leads to more robust models, by reducing the effects of confounding. This work is supported by an NSF CAREER Award, and has garnered attention from other domains where models should agree, at least in part, with domain knowledge. In addition, we have shown how domain knowledge can help in designing network architectures to exploit domain specific-invariances. Invariances are transformations that when applied to the input do not affect the output. Such transformations are often domain or task-specific. E.g., many of the architectures proposed in computer vision are designed to efficiently exploit invariances that arise in tasks involving natural images (e.g., translation invariance). Using domain knowledge about the presence/absence of invariances in tasks involving medical images, we proposed two new CNN architecture modifications tailored to brain images. In contrast to a standard CNN, in which the same filter/feature is convolved over the entire image, our approach learns region-specific features, since a pattern may have a different meaning depending on where in the brain it arises.
Sequence Transformer Networks – When training data are limited, ML practitioners can augment their training data by transforming their data in such a way that invariances are exploited e.g., cropping, flipping, or rotating an image. However, given heterogeneous clinical time-series data, data augmentation is not straightforward. Still, we expect certain temporal invariances to arise in tasks involving clinical time-series data, since oftentimes the relative ordering of events is more important than their precise timing. This intra-class variation can be addressed using techniques like dynamic time warping, but at inference time such algorithms are slow. With the goal of speeding up inference, while exploiting task-specific temporal invariances, we developed techniques for automatically learning invariances. Our proposed approach, sequence transformer networks (STNs), learns to transform clinical time-series data, so as to minimize intra-class variations. Applied to the task of predicting in-hospital mortality, this led to improvements in predictive performance over state-of-the-art.
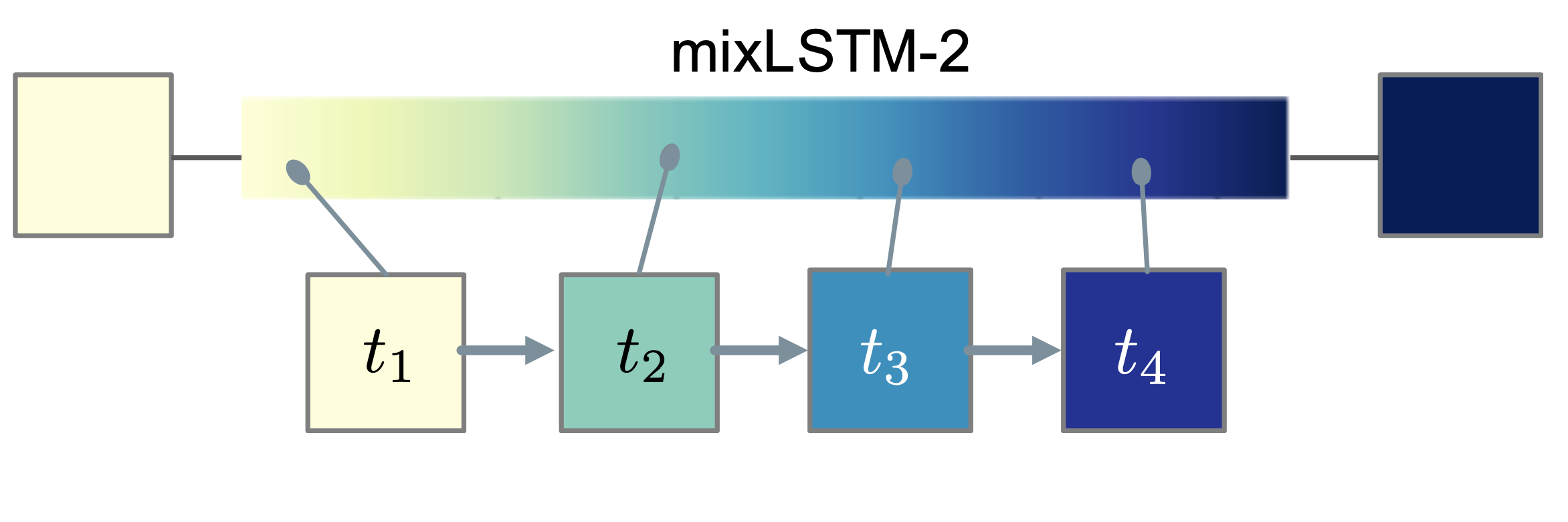
Relaxed Parameter Sharing over Time – In addition, to exploiting invariances, it is important to recognize what invariances are not present in a task. Specifically, we have identified temporal shift in many healthcare tasks, in which the relationship between the input and the output changes over time (e.g., when risk factors change over time). Accurately modeling these time-varying relationships is especially difficult with limited training data. Thus, we recently proposed a new approach for relaxed weight sharing, mixLSTM. Our approach learns multiple sets of parameters and how to combine these parameters at each time step. This is more parameter efficient compared to simply learning different parameters at each time step, and more flexible than a recurrent neural network structure in which parameters are shared across all time steps. Applied to three clinically relevant in-patient prediction tasks, the proposed approach led to significant improvements over several state-of-the-art baselines.
Applications
Learning to Prevent Healthcare-Associated Infections – Healthcare-acquired infections are associated with significant morbidity. In our work on predicting patient risk of infection, We I) pose the problem as a high-dimensional time-series classification task, developing techniques that account for changes over time, and ii) established the importance of hospital-specific models and transfer learning techniques. This work has led to the development of accurate models for identifying patients at risk of C. difficile infection. We have validated the results at six separate hospitals and have shown how a similar approach can be used to predict other outcomes. In addition, we have developed techniques that can account for asymptomatic colonized patients, who may unknowingly spread disease. The work described above culminated in a model tailored to Michigan Medicine (UM) that is currently being applied to daily streams of data to calculate the daily risk of infection of UM inpatients.
Learning from Physiological Signals with Applications in Type I Diabetes – To manage blood glucose levels, individuals with type 1 diabetes (T1D) must constantly make decisions about their regimen. To alleviate this decision fatigue, we are working on novel prediction and control techniques that aim to accurately predict glucose values and the amount of required insulin. More specifically, when learning to predict hyper- and hypoglycemic events, we have shown i) how jointly modeling patterns in the data and the context under which those patterns occur improves performance [KDD’17 ] and ii) that when forecasting blood glucose values, architectures that automatically encode temporal dependencies, while constraining the output, can lead to more accurate forecasts [KDD’18b ]. Currently, we are developing reinforcement learning techniques to learn policies for controlling blood glucose [ICML-WKSP’19 ]. We are now working with JDRF and endocrinologists to translate this work to tools that can improve the lives of people with T1D.

 MENU
MENU